Genomic language model mitigates chimera artifacts in nanopore direct RNA sequencing
Jan 10, 2026·, ,,,,·
1 min read
,,,,·
1 min read
Yangyang Li
Ting-You Wang
Qingxiang (Allen) Guo
Yanan Ren
Xiaotong Lu
Qi Cao
Rendong Yang
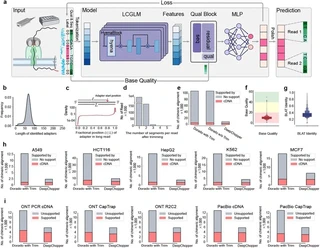
Abstract
Chimera artifacts in nanopore direct RNA sequencing (dRNA-seq) introduce substantial inaccuracies, complicating downstream applications such as transcript annotation and gene fusion detection. Current basecalling models are unable to detect or mitigate these artifacts, limiting the reliability and utility of dRNA-seq for transcriptomics research. To address this challenge, we present DeepChopper, a genomic language model specifically designed to identify and remove adapter sequences from base-called dRNA-seq long reads with single-base precision. Operating independently of raw signal or alignment information, DeepChopper effectively eliminates adapter-bridged artifacts. Here, we show that DeepChopper enhances the accuracy of downstream analyses and unlocks the full potential of nanopore dRNA-seq, establishing it as a more robust tool for diverse transcriptomics applications.
Type
Publication
Nature Communications
Official Publication: https://doi.org/10.1038/s41467-026-68571-5
Preprint (bioRxiv): https://doi.org/10.1101/2024.10.23.619929
Authors
Authors

Authors
Qingxiang (Allen) Guo
(he/him)
Postdoctoral Scholar | Cancer Genomics & AI
Postdoctoral scholar at Northwestern University developing computational approaches for cancer genomics.
My research integrates long-read sequencing, structural variant analysis, and deep learning to decode
the regulatory complexity of cancer genomes.
Authors
Authors
Authors
Authors