I am a computational biologist and postdoctoral fellow in Dr. Rendong Yang’s lab at the Feinberg School of Medicine, Northwestern University. My research focuses on developing advanced bioinformatics algorithms, deep learning models, and multi-omic pipelines to decode the regulatory complexity of cancer genomes and transcriptomes. I am particularly passionate about leveraging third-generation long-read sequencing (e.g., Oxford Nanopore) and AI-driven methods to uncover novel structural variations and transcriptomic alterations that traditional methods often miss.
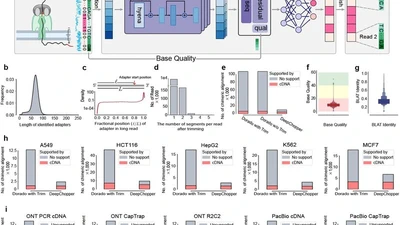
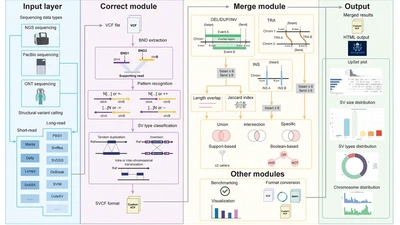
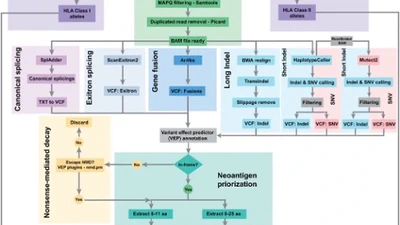
Currently, my work centers on building computational tools that address critical bottlenecks in biomedical genomics. As a lead developer, I created toolkits like OctopuSV (Bioinformatics) for automated, multi-sample structural variant analysis. Furthermore, I have enjoyed collaborating with excellent colleagues on various impactful projects. For instance, I contributed to the project of DeepChopper—a genomic language model published in Nature Communications designed to mitigate chimera artifacts in nanopore direct RNA sequencing. Additionally, I participated in developing analytical workflows for neoantigen detection (ScanNeo2, Bioinformatics) and supported the analysis for discovering novel cancer regulatory mechanisms (Science Advances).
My doctoral training provided a rigorous methodological foundation in algorithm design and multi-omic integration, which I further refined through independent computational development and cross-disciplinary collaborations. My scientific perspective was shaped during my studies at Oregon State University in 2014. I am grateful to Prof. Jerri Bartholomew for her mentorship, which sparked my interest in evolutionary genomics and laid the groundwork for my current research. To date, I have authored and co-authored 26 peer-reviewed publications, including works published in Nature Communications, Science Advances, Bioinformatics, BMC Biology, and Biology (eight papers as first author).
My long-term goal is to become an independent academic investigator advancing the intersection of AI and biomedical genomics. By designing specialized long-read and deep learning-based tools, I aim to bridge the gap between computational innovation and clinical application—ultimately enabling earlier cancer detection, precise target discovery, and the development of personalized therapies.